Force atlas in cytoscape11/5/2023 Analysis can be performed via the Cytoscape GUI or CyREST programming interface using R (RC圓) or Python (py4cytoscape).Īpp Cytoscape Expression analysis Network biology Single cell scRNA-seq. scNetViz enables users to analyze data from public atlases or their own experiments, which we illustrate with two use cases. We describe our implementation of methods for accessing data from public single cell atlas projects, differential expression analysis, visualization, and automation. 
To automate a complete data analysis workflow, scNetViz integrates parts of the Scanpy software, which is a popular Python package for scRNA-seq data analysis, with Cytoscape apps such as stringApp, cyPlot, and enhancedGraphics. scNetViz calculates the differential expression of each gene across clusters and then creates a cluster-specific gene functional interaction network between the significantly differentially expressed genes for further analysis, such as pathway enrichment analysis. Here, we present scNetViz - a Cytoscape app to aid biological interpretation of cell clusters in scRNA-seq data using network analysis. Users can adjust the force between nodes in a given set relative to connected nodes and thereby emphasize or diminish the grouping based on set membership by changing. 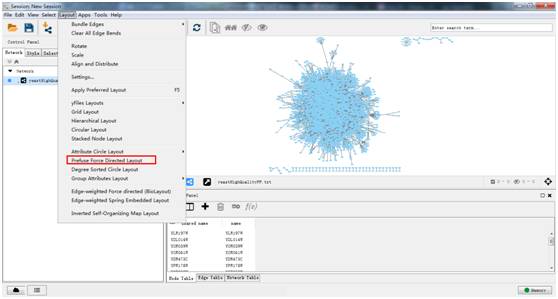 
Applying network biology approaches to scRNA-seq data can provide useful insights into genes driving heterogeneous cell-type compositions of tissues. The setsbased force directed layout employs the Prefuse force directed layout provided by Cytoscape but also tries to put nodes in the same set closer together in the network. Single-cell RNA-sequencing (scRNA-seq) has revolutionized molecular biology and medicine by enabling high-throughput studies of cellular heterogeneity in diverse tissues.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
